Should I Use Dask?

At Saturn, we love Dask, but we still recommend maximizing simpler approaches before using it. This article gives an overview of good reasons to choose Dask, as well as alternatives that should be explored before you take the plunge. Runtime speed is a big blocker for data science iteration speed, which is ultimately a blocker for the business value you hope to get from your team of data scientists. No matter what approach you take, maximizing iteration speed of your data scientists is the biggest value add you can make.
What is Dask?
From dask.org, “Dask is a flexible library for parallel computing in Python”. For most users, Dask is 3 things:
- A pandas-like DataFrame, which can take advantage of multiple processors or machines.
- A numpy-like Array, which can take advantage of multiple processes or machines.
- A way to write (delayed) functions, that can run on multiple processes or machines.
Dask is pretty great, but it also adds complexity.
Dask DataFrame
A Dask DataFrame is represented by many smaller Pandas DataFrames that live on your cluster. When you interact with the Dask Dataframe, the functions and methods you call are converted into many smaller functions which are executed in parallel on the smaller Pandas DataFrames.
Additional resources:
- How to use Dask DataFrames as an alternative to pandas across multiple machines
- Dask DataFrame is not Pandas
- Getting Started With Dask
Dask Array
A Dask Array is represented by many smaller NumPy Arrays that live on your cluster. When you interact with the Dask Array, the functions and methods you call are converted into many smaller functions that are executed in parallel on the smaller NumPy Arrays.
Dask Delayed
A Dask delayed function is a function that is designed to run on a Dask cluster. When you call a delayed function, instead of evaluating the function, you get a “delayed” result which represents the future result of that computation.
Running Dask?
Most people start out running Dask on a single machine, with multiple worker threads or processes serving as your cluster. This is called the Dask LocalCluster. For bigger problems, people generally run many dask workers on a cluster of multiple machines. Setting this up in a secure and useful way isn’t straightforward, and that’s why we exist!
Should I use Dask?
There area few good reasons to do so:
- Are you running out of memory on your workstation?
- Are you waiting more than 20 minutes for your work to run?
- Do you need more GPUs?
Solving memory issues without Dask
If you’re running out of memory on your workstation, Dask can help, but there are a few things I would recommend doing first. If you’re using primarily NumPy arrays, you can skip this section. Other than moving to sparse arrays (which have their own problems), there isn’t much you can do to optimize memory in NumPy. If you’re using Pandas, here are some suggestions.
StringDType
Python strings have roughly 40 bytes of overhead. That doesn’t sound
like a lot, but if you have a billion strings, it can add up quickly.
The new StringDType can help here.
df['column'] = df['column'].astype(pd.StringDType())
Categorical DTypes
Many string and numerical fields are actually categorical. Take a column named “Household Income”. Instead of a numerical value, you usually get data in bands, like “0-$40,000” or “more than $100,000”.
df['column'] = df['column'].astype('category')
As a general guideline, I usually look for columns where the ratio of the number of unique values, to the number of rows is less than 1%.
Diagnosing Memory Usage
df.memory_usage(deep=True) is a good
way to understand how much memory each column is using. This can help
you understand what benefit you are getting from converting your data to
appropriate DTypes.
Get a bigger computer?
If you’re working on the cloud, the easiest solution is usually to get the biggest machine you can. At Saturn, we currently top out at 512 GB (but let us know if you want more RAM!)
Dask?
If you’re done optimizing memory, and your dataset takes up
approximately 1/3 of your workstations memory, I would recommend
skipping Dask. If your dataset takes up more than 1/3 of your
workstations memory, there is a good chance that subsequent operations
you do on that data frame will exceed the memory of your workstation,
and looking at dask.dataframe is a
good solution. If you’re doing this for memory-bound reasons, you’re
going to need a multi-node Dask cluster in order to get more
memory.
Shameless Plug
At Saturn Cloud, we make it really easy to get a beefy Jupyter server where you can run a Dask LocalCluster. We also make it really easy to start a multinode cluster.
cluster = SaturnCluster(worker_size="g4dnxlarge", scheduler_size="medium", nprocs=1, nthreads=4)
client = Client(cluster)
if you’re running into any of these problems and you want to try Dask, give us a whirl!
Solving compute time issues without Dask
If you’ve got some code that’s slow, Dask can help! But there are a few things to check first.
Swap Pandas for NumPy
NumPy can be orders of magnitude faster for many operations compared to
Pandas. This is the biggest problem in a piece of code that gets called
many times, for example, the objective function to any call
to scipy.optimize. Replacing Pandas DataFrame calls with NumPy calls can
easily speed up your code by 50x
Profile your code
There are many Python profiling tools, but the line_profiler is the
easiest to understand. Profiling is a long topic, the short version here
is, once you have line_profiler installed, you can call
%load_ext line_profiler
To load it into Jupyter, and then execute:
%lprun -f func1 -f func2 expression
The lprun command takes multiple -f parameters, which identify the functions you want to profile. The expression is then evaluated, and the profile results displayed.
For example, if I have functions test,
and test2, I might execute
%lprun -f test -f test2 test()
You will then get an output like:
#> Line # Hits Time Per Hit % Time Line Contents
#> ==============================================================
#> 149 @profile150 def Proc2(IntParIO):
#> 151 50000 82003 1.6 13.5 IntLoc = IntParIO + 10
#> 152 50000 63162 1.3 10.4 while 1:
#> 153 50000 69065 1.4 11.4 if Char1Glob == 'A':
#> 154 50000 66354 1.3 10.9 IntLoc = IntLoc - 1
#> 155 50000 67263 1.3 11.1 IntParIO = IntLoc - IntGlob
#> 156 50000 65494 1.3 10.8 EnumLoc = Ident1
#> 157 50000 68001 1.4 11.2 if EnumLoc == Ident1:
#> 158 50000 63739 1.3 10.5 break
#> 159 50000 61575 1.2 10.1 return IntParIO
Which Dask?
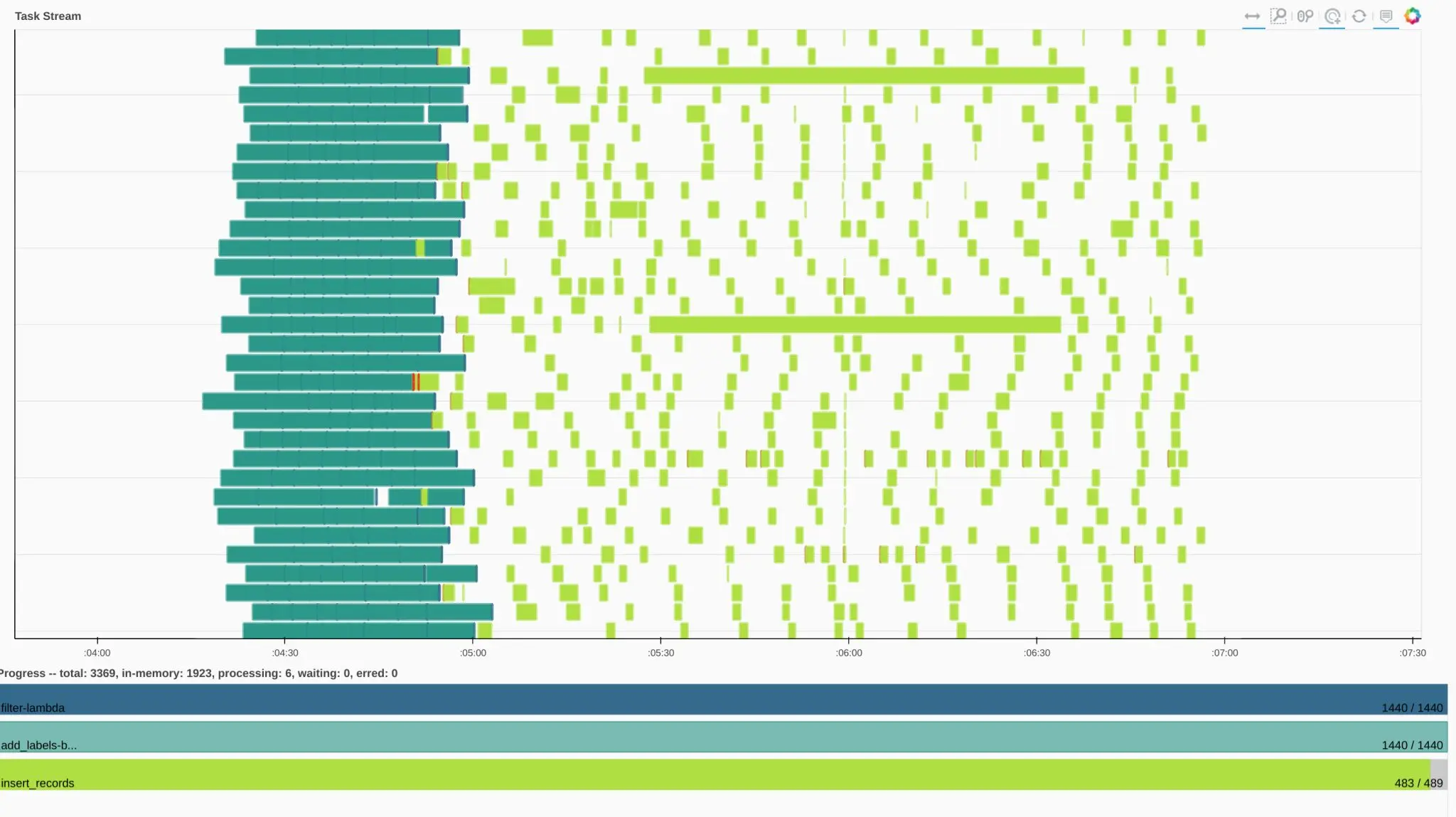
I’d start with the least amount of complexity – max out the number of cores on your laptop. You can do this with multiprocessing, however I find single node Dask to be easier to use, and more convenient than multiprocessing. The Dask dashboard is also extremely helpful in helping you understand what’s happening on your machine.
But I need more GPUs
Short of shoving more GPU cards into your machine, there isn’t a good way to solve this problem without going multi-node. This is a good problem to solve with Dask. Assuming you’re already running in the cloud, the biggest GPU instance on AWS has 8 v100 GPUs. If you’re going to need more of them, using Dask to build a cluster of v100 nodes is a good option.
How do I use Dask
This obviously deserves its own article, or series of articles, but let’s cover it very briefly.
DataFrame and Array
Dask DataFrame has a bunch of methods like read_csv,
or read_parquet, That can read a collection of data files, and return a
distributed DataFrame. In the most general case, if you have a function
that can return a small subset of your data, you can do something like
this
from dask import delayed
import dask.dataframe as dd
@delayed
def get_partition(partition_number):
...
partitions = [get_partition(num) for num in range(number_of_partitions)]
df = dd.from_delayed(partitions)
A similar approach works for Dask Array
Delayed
from dask import delayed
@delayed
def inc1(x):
return x + 1
@delayed
def sum(x):
return sum(x)
result = sum([inc1(i) for i in input_vars])
Here, result represents a delayed computation. Nothing has been executed yet, until:
result = result.compute()
Conclusion
Distributed computing is hard. Dask is the easiest way to do it. Saturn Cloud is the easiest way to Dask. If you’re interested, let us know. We regularly do POCs to help people figure out how to get value out of their Dask cluster and often realize 100x speedups. If you’re used to waking up in the middle of the night to make sure your 24 hour long python job hasn’t crashed, it’s time to live your best life.
We offer several plans that meet individual, team, and enterprise needs.
About Saturn Cloud
Saturn Cloud is your all-in-one solution for data science & ML development, deployment, and data pipelines in the cloud. Spin up a notebook with 4TB of RAM, add a GPU, connect to a distributed cluster of workers, and more. Request a demo today to learn more.



